Use the Off-Spotter web tool | Learn how to use Off-Spotter
If you use any of the below please cite: Pliatsika, V, and Rigoutsos, I (2015) Off-Spotter: very fast and exhaustive enumeration of genomic lookalikes for designing CRISPR/Cas guide RNAs Biol. Direct 10(1):4
In this page you can find the source code of Off-Spotter1. Off-Spotter is our new tool that identifies all genome hits of one or multiple gRNAs. Given a sequence as input, it returns the gRNAs together with position, structure, and annotation information. Off-Spotter is extremely fast and accurate.
Off-Spotter is also very flexible. You can use your own genome data and any number of annotation files. It is set up to work for 4 PAM sequences, NGG, NAG, NNNNACA, and NNGRRT (where R is A or G). Upon completion it returns the chromosome number, strand, starting position, ending position, the input gRNA, the hit, the PAM, the number of mismatches, and annotation information. To learn more about Off-Spotter visit our Off-Spotter help page.
Web Tool
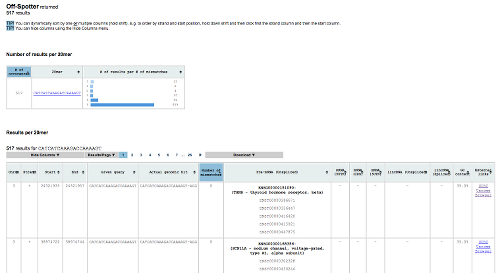
In addition to source code, Off-Spotter is also available as a web tool2. The web tool provides all the information that the source code provides. Additionally it provides numerous sorting and viewing options as well as statistic information for the hits. Click here to use our web tool.
Downloads
Off-Spotter Source code
Current version: 0.2.2, released 10/26/2015
You can download the Off-Spotter source code as a single compressed file by clicking below. By downloading, you agree on our terms and conditions. See the changelog. For more information on how to use the code please visit our help page.
Unsupported older version:
Off-Spotter Remote submission tool
Current Version: 1.2, released 3/22/2022
You can also download this utility that enables you to submit data to Off-Spotter from your computer. Use this for automated queries, larger inputs, or several sequences.
Terms and Conditions
This code (© 2015 Thomas Jefferson University, All Rights Reserved) was created by Venetia Pliatsika and Isidore Rigoutsos and is an implementation of the Off-Spotter algorithm that appears in Pliatsika, V, and Rigoutsos, I (2015) “Off-Spotter: very fast and exhaustive enumeration of genomic lookalikes for designing CRISPR/Cas guide RNAs” Biol. Direct 10(1):4.
Use of these codes is bound by the following terms and conditions:
Terms of Use:
This code can be freely used for research, academic and other non-profit activities. Only one instance of the code may be used at a time, and then for only one concurrent user. You may not use the code to conduct any type of application service, service bureau or time-sharing operation or to provide any remote processing, network processing, network telecommunications or similar services to any person, entity or organization, whether on a fee basis or otherwise. The code can be copied and compiled on any platform for the use authorized by these terms and conditions. All copies of the code must be accompanied by this note. The code cannot be modified without the written permission of the Computational Medicine Center of Thomas Jefferson University https://cm.jefferson.edu
Commercial use is strictly prohibited. If you wish to use these codes commercially please contact the Computational Medicine Center of Thomas Jefferson University: https://cm.jefferson.edu/contact-us/
THE CODE IS PROVIDED “AS IS” WITH NO REPRESENTATIONS OR WARRANTIES OF ANY KIND, EITHER EXPRESSED OR IMPLIED. TO THE FULLEST EXTENT PERMISSIBLE PURSUANT TO APPLICABLE LAW. THOMAS JEFFERSON UNIVERSITY, AND ITS AFFILIATES, DISCLAIM ALL WARRANTIES, EXPRESS OR IMPLIED, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF TITLE, MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NON-INFRINGEMENT.
NEITHER THOMAS JEFFERSON UNIVERSITY NOR ITS AFFILIATES MAKE ANY REPRESENTATION AS TO THE RESULTS TO BE OBTAINED FROM USE OF THE CODE.
References
- Pliatsika, V, and Rigoutsos, I (2015) Off-Spotter: very fast and exhaustive enumeration of genomic lookalikes for designing CRISPR/Cas guide RNAs Biol. Direct 10(1):4
- Off-Spotter web tool
Contact us for licensing inquiries or to report bugs.