Use the Teiresias webapp | Learn more about Teiresias
If you use any of the below please cite: Rigoutsos, I, Floratos, A (1998) Combinatorial pattern discovery in biological sequences: The TEIRESIAS algorithm. Bioinformatics 14: 55-67
The Teiresias algorithm is a combinatorial algorithm for the discovery of rigid patterns (motifs) in biological sequences. It is named after the Greek prophet Teiresias and was created in 1997 by Isidore Rigoutsos and Aris Floratos. Teiresias should be used for the discovery of exact or more lenient similarities within a set of character sequences […] read more on the “how to use Teiresias” page.
Web Tool
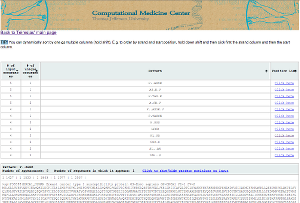
Teiresias is also available as a web tool. You can access the web tool and use Teiresias online here. For more information as to how to use the web tool, please click here.
Tutorial and version information
For more details on how to use Teiresias please visit the “how to use Teiresias” page.
Terms and Conditions
By downloading these files you are agreeing to the following terms and conditions.
Terms of Use:
This code can be freely used for research, academic and other non-profit activities (the “Authorized Use”). Commercial use is strictly prohibited. The code can be copied and compiled on any platform for the Authorized Use, but cannot be modified without the written permission of the Computational Medicine Center of Thomas Jefferson University. THE CODE IS PROVIDED “AS IS” WITH NO REPRESENTATIONS OR WARRANTIES OF ANY KIND, EITHER EXPRESSED OR IMPLIED. TO THE FULLEST EXTENT PERMISSIBLE PURSUANT TO APPLICABLE LAW. THOMAS JEFFERSON UNIVERSITY, AND ITS AFFILIATES, DISCLAIM ALL WARRANTIES, EXPRESS OR IMPLIED, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF TITLE, MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NON-INFRINGEMENT. NEITHER THOMAS JEFFERSON UNIVERSITY NOR ITS AFFILIATES MAKE ANY REPRESENTATION AS TO THE RESULTS TO BE OBTAINED FROM USE OF THE CODE.Download
You can download the Teiresias source files as a single compressed file by clicking below.
Current version: 0.9.1 RELEASED 3/13/2014
References
- Rigoutsos, I, Floratos, A (1998) Combinatorial pattern discovery in biological sequences: The TEIRESIAS algorithm. Bioinformatics 14: 55-67
For any concerns or to report bugs, you can contact us here
