Pancreatic cancer is among the cancer types with the worst prognosis. The TCGA network performed a multi-platform analysis of 150 tumors. A cofounding factor was the neoplastic cellularity (or tumor purity), the relative ratios of the whole tumor in cancer cells vs. stromal cells. As part of working group, we devised a two-level analysis working on the most pure tumors separately from the samples with lower tumor purity. This strategy proved successful in all of the levels of cellular functions analyzed, from genomic analysis to phosphor-proteomics.
Our data-driven and hypothesis-generating analyses utilized transcriptomic and methylation data and focused on expanding and integrating the mRNA-defined subtypes with the miRNA and methylation level. Specifically, as the TCGA mRNA data provide a deep and comprehensive analysis of the pancreatic cancer subtypes and their differences, we wanted to examine how these differences emerge and how could they be regulated epigenetically, by genome methylation or miRNAs. We were excited to find two miRNAs being very central in this process, miR-192 and miR-194. They were negatively correlated with several mRNAs, the corresponding genes and gene products of which are the ones responsible for the two major pancreatic cancer subtypes. Interestingly, the miRNA levels were opposite of methylation levels, indicating an important epigenetic regulation, a different “state” that is not visible with conventional mutation analyses.
We also found an opportunity to use the not-pure samples to extract an mRNA-miRNA interaction signature that characterizes the stromal cells and not the cancer cells. A very important miRNA, miR-21 emerged in these analyses, suggesting that its role is potentially more important in the tumor stroma than in cancer, at least in pancreatic cancer.
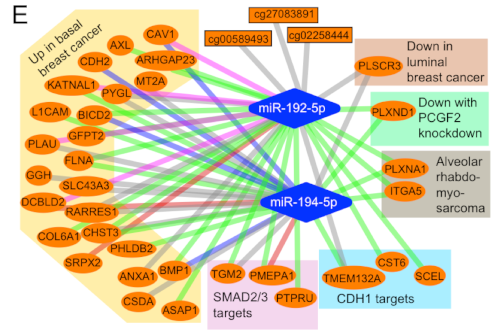 Role of miR-192-5p and miR-194-5p in cancer
Role of miR-192-5p and miR-194-5p in cancer
You can read more on the TCGA paper, published in Cancer Cell.
References
- Cancer Genome Atlas Research Network. Electronic address: andrew_aguirre@dfci.harvard.edu, Cancer Genome Atlas Research Network. Integrated Genomic Characterization of Pancreatic Ductal Adenocarcinoma. Cancer Cell. 2017;32 (2):185-203.e13. doi: 10.1016/j.ccell.2017.07.007. PubMed PMID:28810144 PubMed Central PMC5964983.
