Eukaryotic cells equip robust systems to respond to stress conditions. In stressed mammalian cells, angiogenin endoribonuclease cleaves anticodon-loops of tRNAs to generate tRNA halves termed tRNA-derived stress-induced RNAs (tiRNAs), which promote stress granule formation and regulate translation. While the molecular functions of tiRNAs have been demonstrated, the expression profile of tiRNAs remains elusive. The 5′-tiRNAs (5′-tRNA halves) contain a 2′,3′-cyclic phosphate (cP) and thus belong to cP-containing RNAs (cP-RNAs). The cP-RNAs form a hidden layer of the transcriptome because standard RNA-seq cannot amplify and sequence them.
In this study, we performed genome-wide analyses of short cP-RNA transcriptome in oxidative stress-exposed human cells. Using cP-RNA-seq that can specifically sequence cP-RNAs, we identified tiRNAs and numerous other cP-RNAs that are mainly derived from rRNAs and mRNAs. Although tiRNAs were produced from a wide variety of tRNA species, abundant species of tiRNAs were derived from a focal-specific subset of tRNAs. Regarding rRNA- and mRNA-derived cP-RNAs, determination of the processing sites of substrate RNAs revealed highly specific RNA cleavage events between pyrimidines and adenosine in generation of those cP-RNAs. Those cP-RNAs were derived from specific loci of substrate RNAs rather than from the overall region, implying that cP-RNAs are produced by regulated biogenesis pathways and not by random degradation events. We experimentally confirmed the identified sequences to be expressed as cP-RNAs in the cells, and their expressions were upregulated upon induction of oxidative stress. These analyses of the cP-RNA transcriptome unravel an abundant class of short ncRNAs that accumulate in cells under oxidative stress.
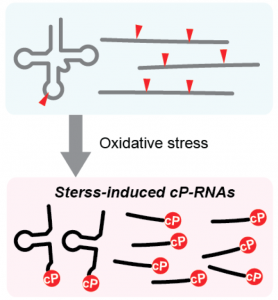 Identification of stress-induced cP-RNAs
Identification of stress-induced cP-RNAs
References
- Shigematsu, M, Kirino, Y. Oxidative stress enhances the expression of 2′,3′-cyclic phosphate-containing RNAs. RNA Biol. 2020. doi: 10.1080/15476286.2020.1766861. PubMed PMID:32397797.
