Transfer RNAs (tRNAs) function in translational machinery and further serves as a source of short non-coding RNAs (ncRNAs). tRNA-derived ncRNAs show differential expression profiles and play roles in many biological processes beyond translation. Molecular mechanisms that shape and regulate their expression profiles are largely unknown. Here we report the mechanism of biogenesis for tRNA-derived Piwi-interacting RNAs (td-piRNAs) expressed in Bombyx BmN4 cells. In the cells, two cytoplasmic tRNA species, tRNAAspGUC and tRNAHisGUG, served as major sources for td-piRNAs, which were derived from the 5′-part of the respective tRNAs. cP-RNA-seq identified the two tRNAs as major substrates for the 5′-tRNA halves as well, suggesting a previously uncharacterized link between 5′-tRNA halves and td-piRNAs. An increase in levels of the 5′-tRNA halves, induced by BmNSun2 knockdown, enhanced the td-piRNA expression levels without quantitative change in mature tRNAs, indicating that 5′-tRNA halves, not mature tRNAs, are the direct precursors for td-piRNAs. For the generation of tRNAHisGUG-derived piRNAs, BmThg1l-mediated nucleotide addition to −1 position of tRNAHisGUG was required, revealing an important function of BmThg1l in piRNA biogenesis. Our study advances the understanding of biogenesis mechanisms and the genesis of specific expression profiles for tRNA-derived ncRNAs.
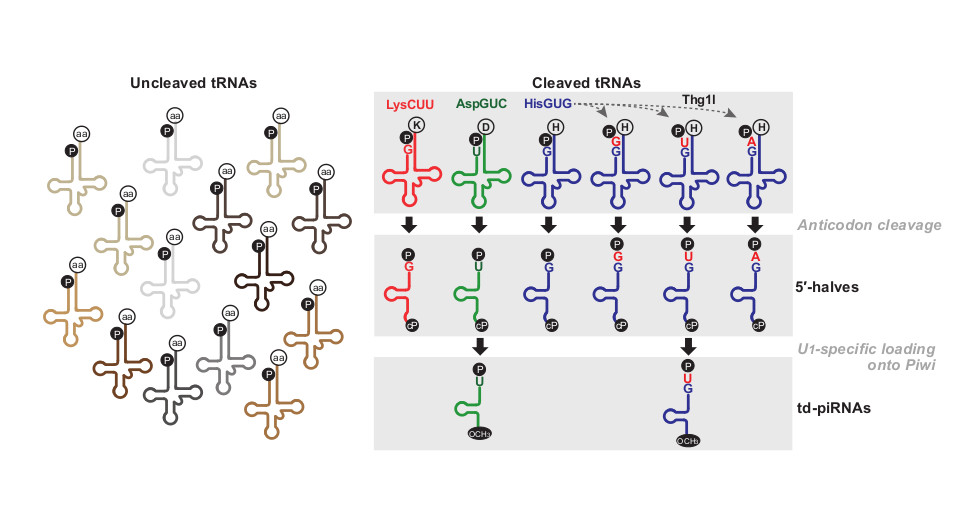 The Biogenesis Pathway of tRNA-Derived piRNAs
The Biogenesis Pathway of tRNA-Derived piRNAs
References
- Honda, S, Kawamura, T, Loher, P, Morichika, K, Rigoutsos, I, Kirino, Y. The biogenesis pathway of tRNA-derived piRNAs in Bombyx germ cells. Nucleic Acids Res. 2017;45 (15):9108-9120. doi: 10.1093/nar/gkx537. PubMed PMID:28645172 PubMed Central PMC5587776.
